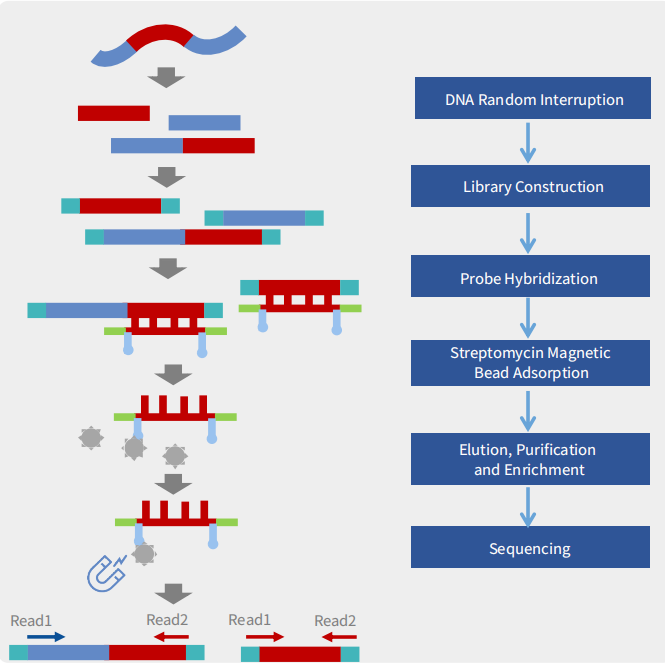
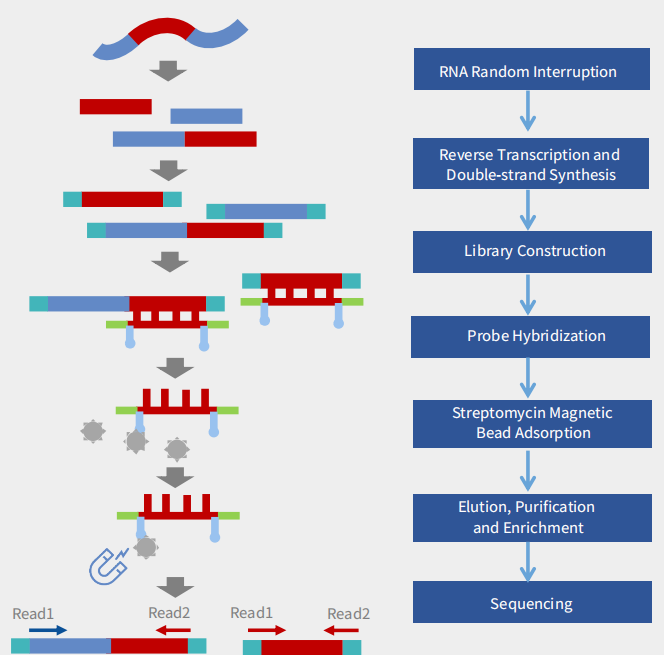
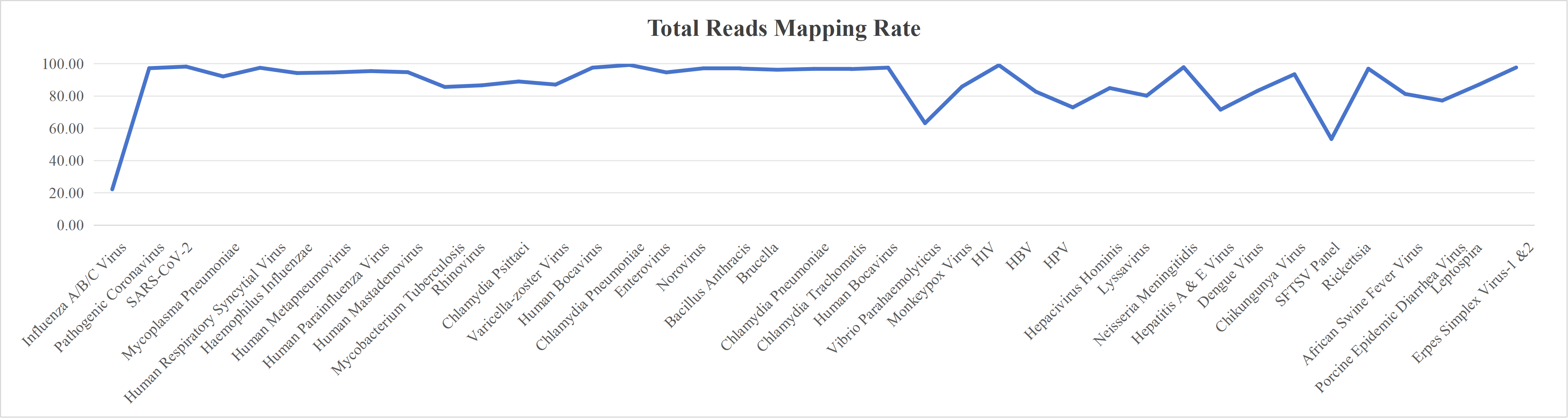
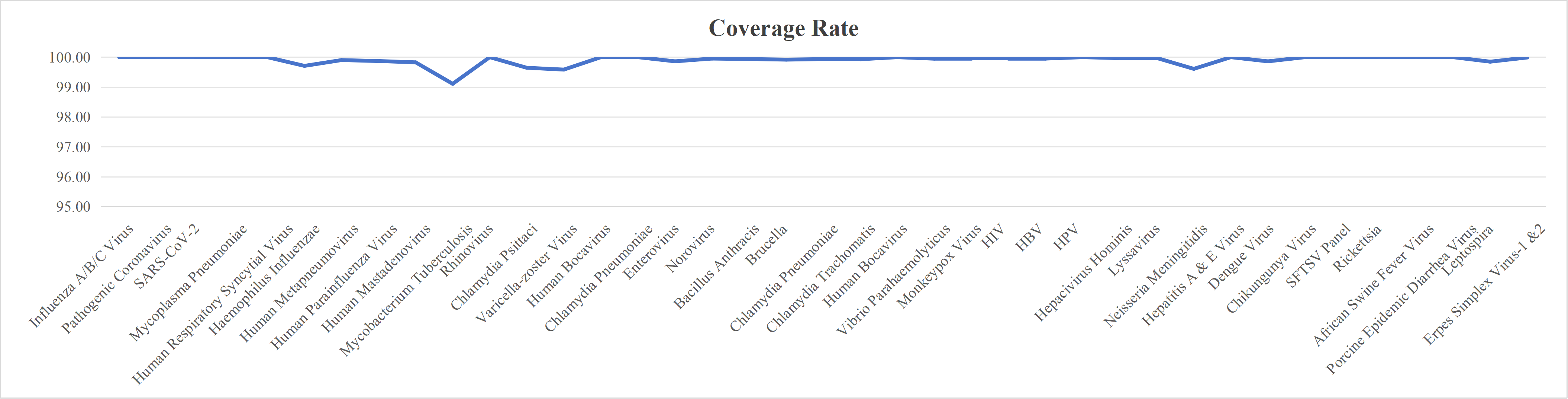
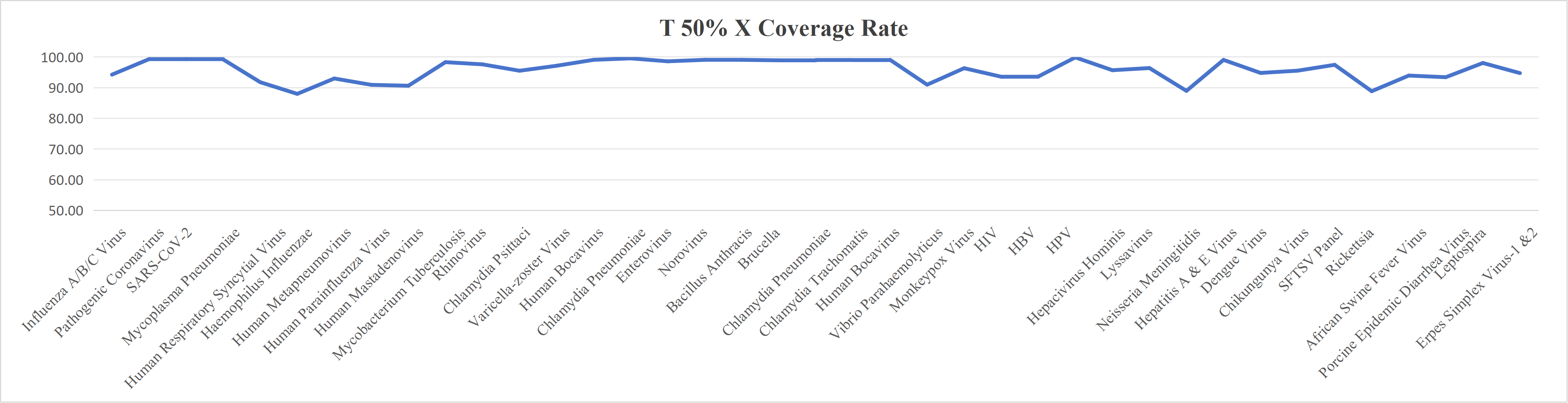
Probe Design
We design high-coverage probes based on all polymorphic sequences of the target pathogen. Using sequence-similarity analysis, redundant probes in conserved regions are removed while preserving probe diversity in highly variable regions, ensuring comprehensive coverage with controlled cost.