Performance — Measured Results
Selected measured metrics and coverage demonstration for representative pathogens (figures from internal validation).
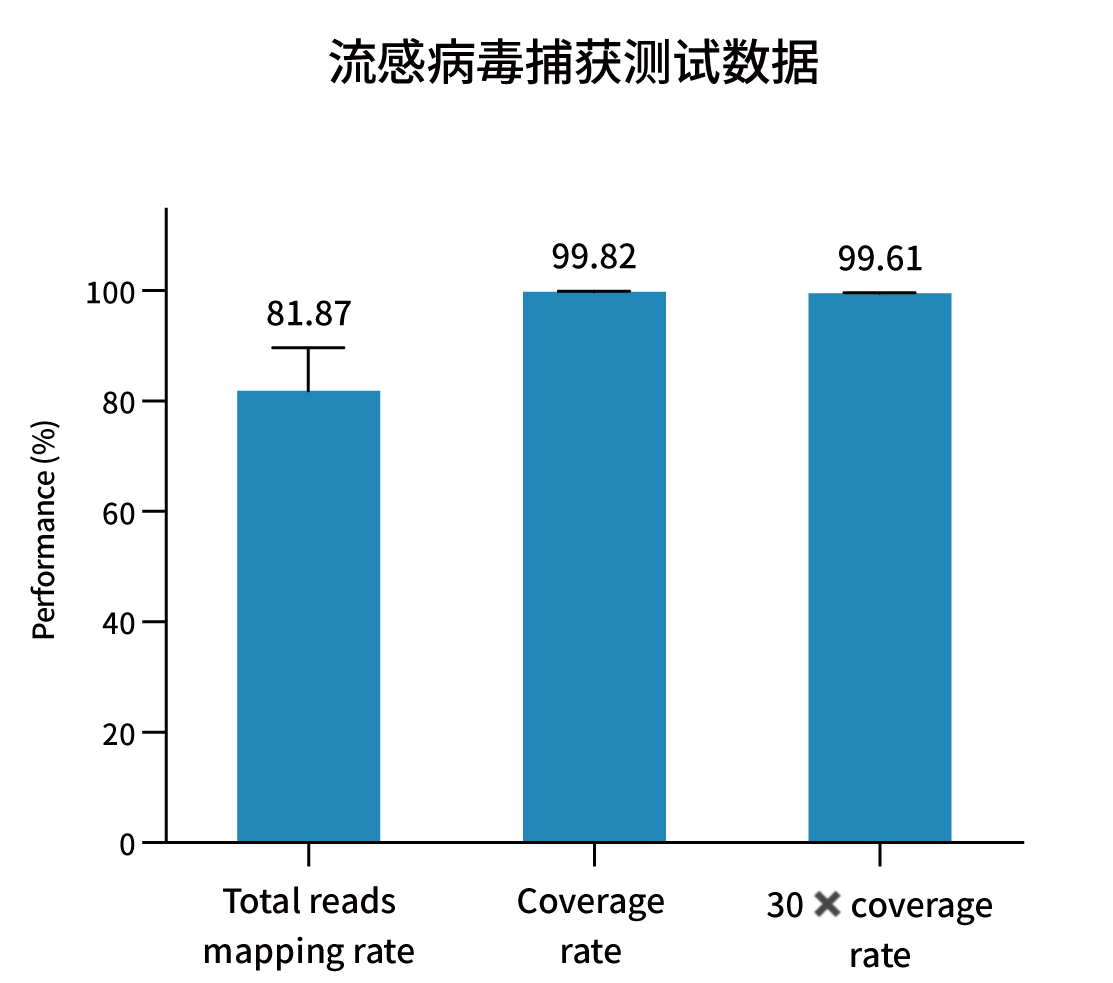
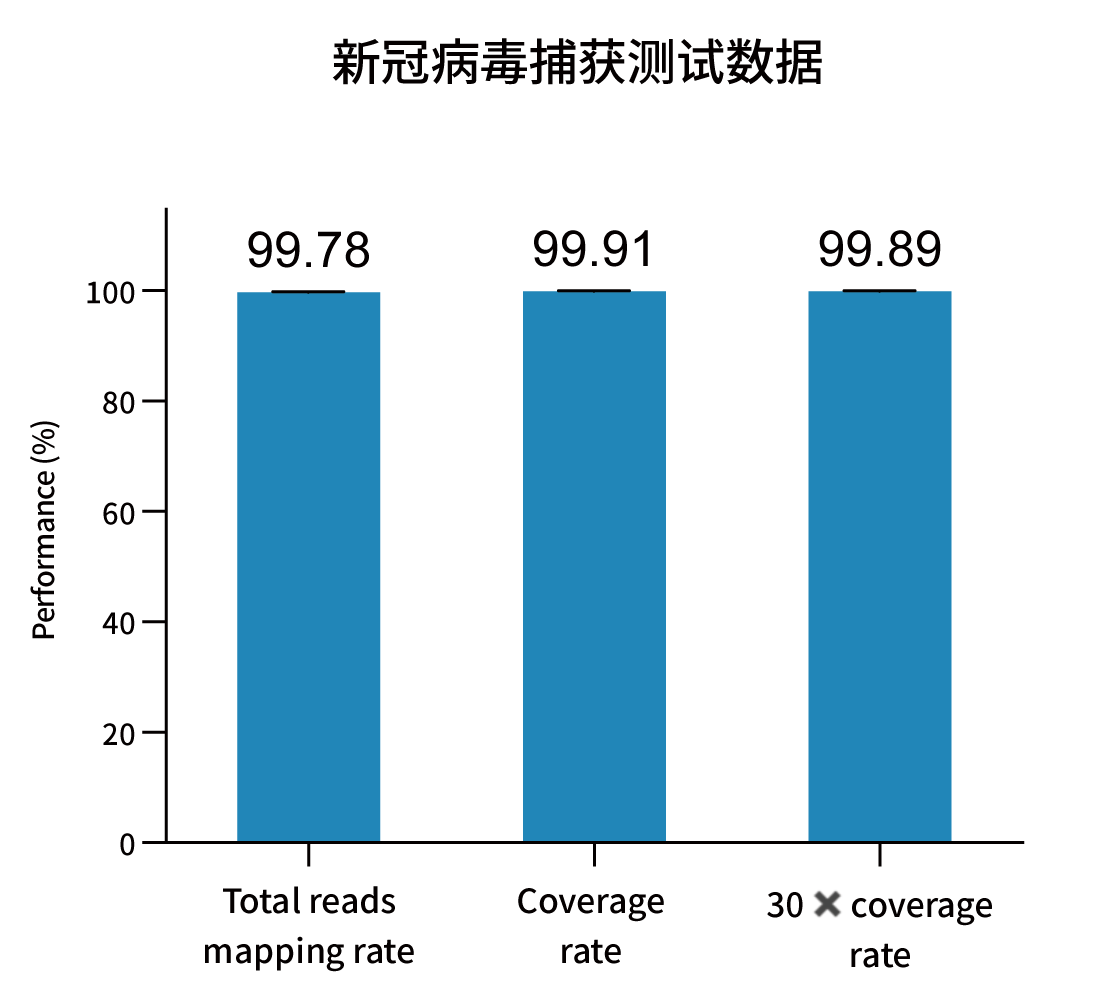
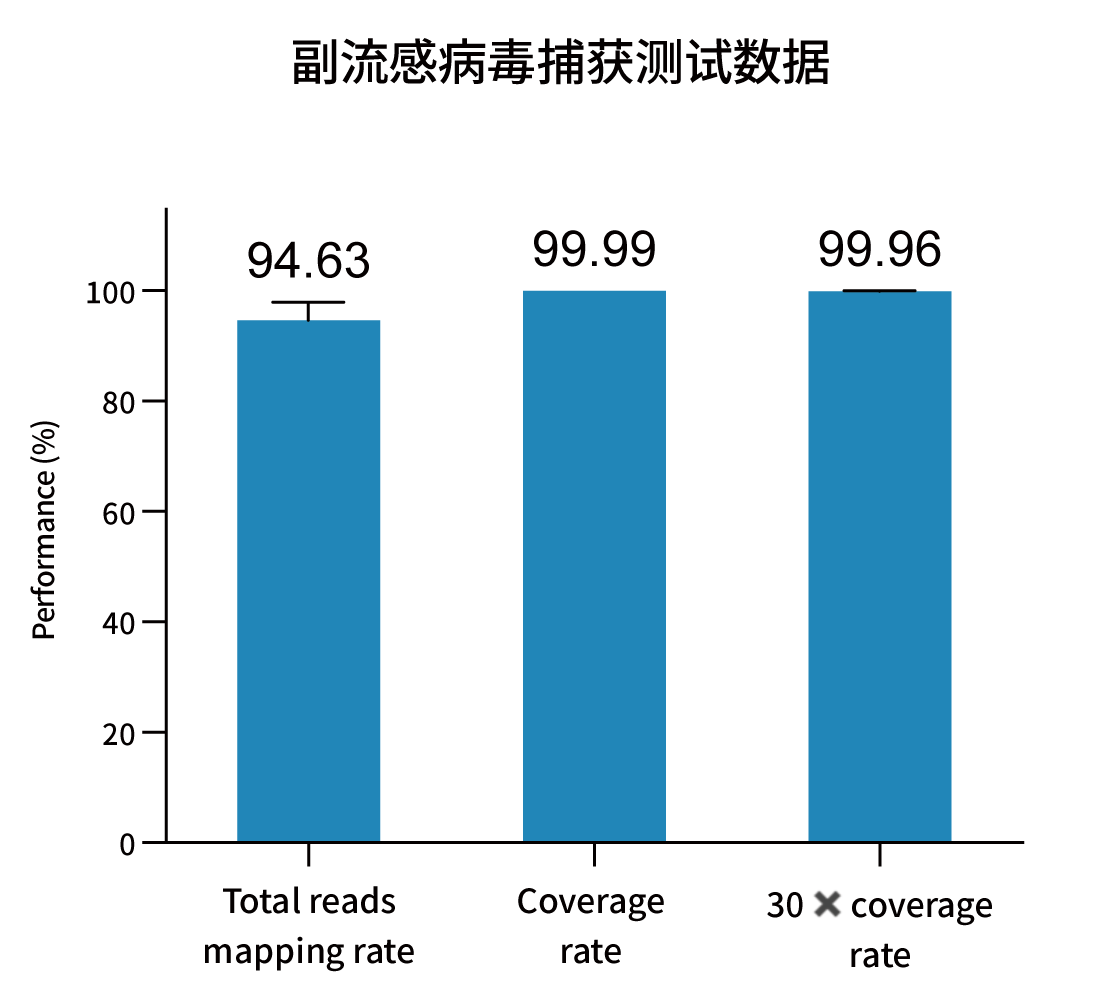
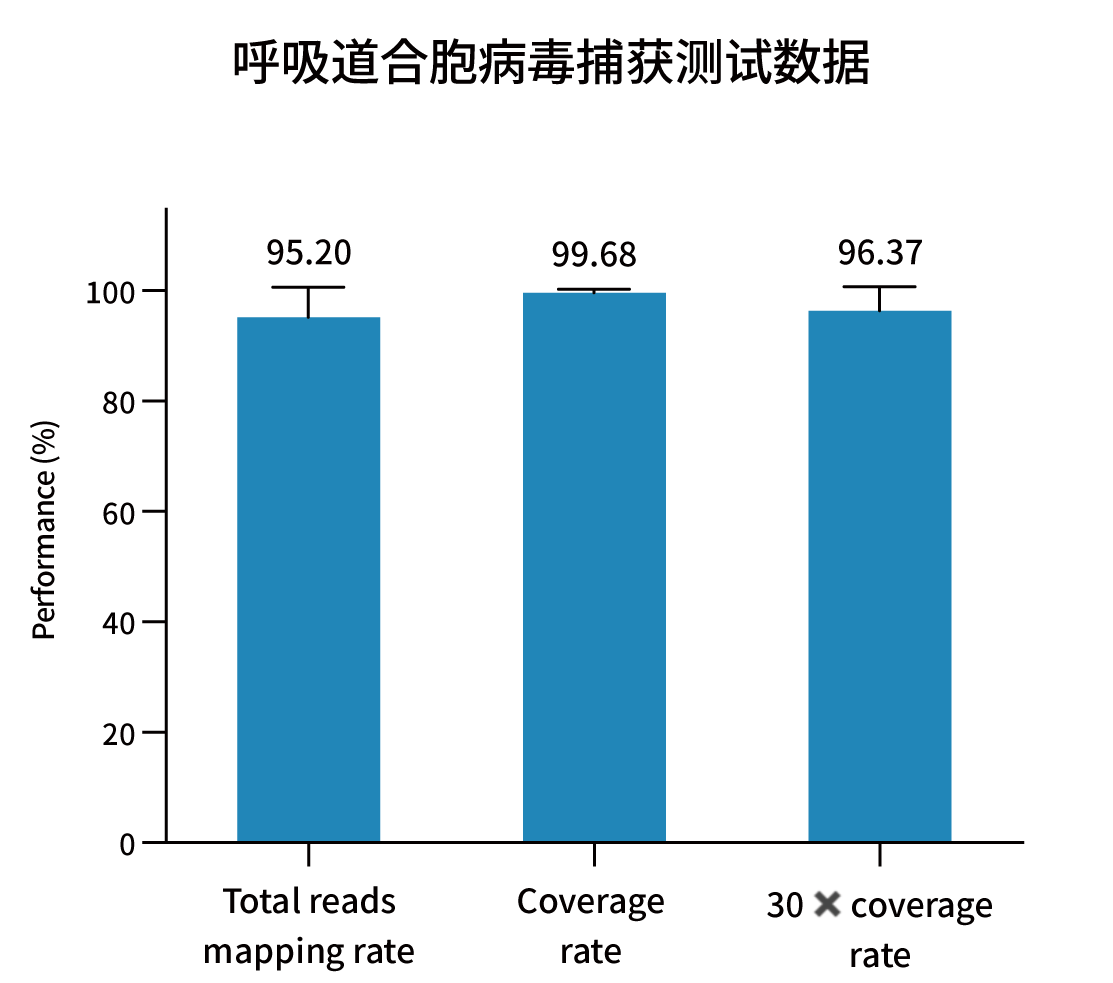
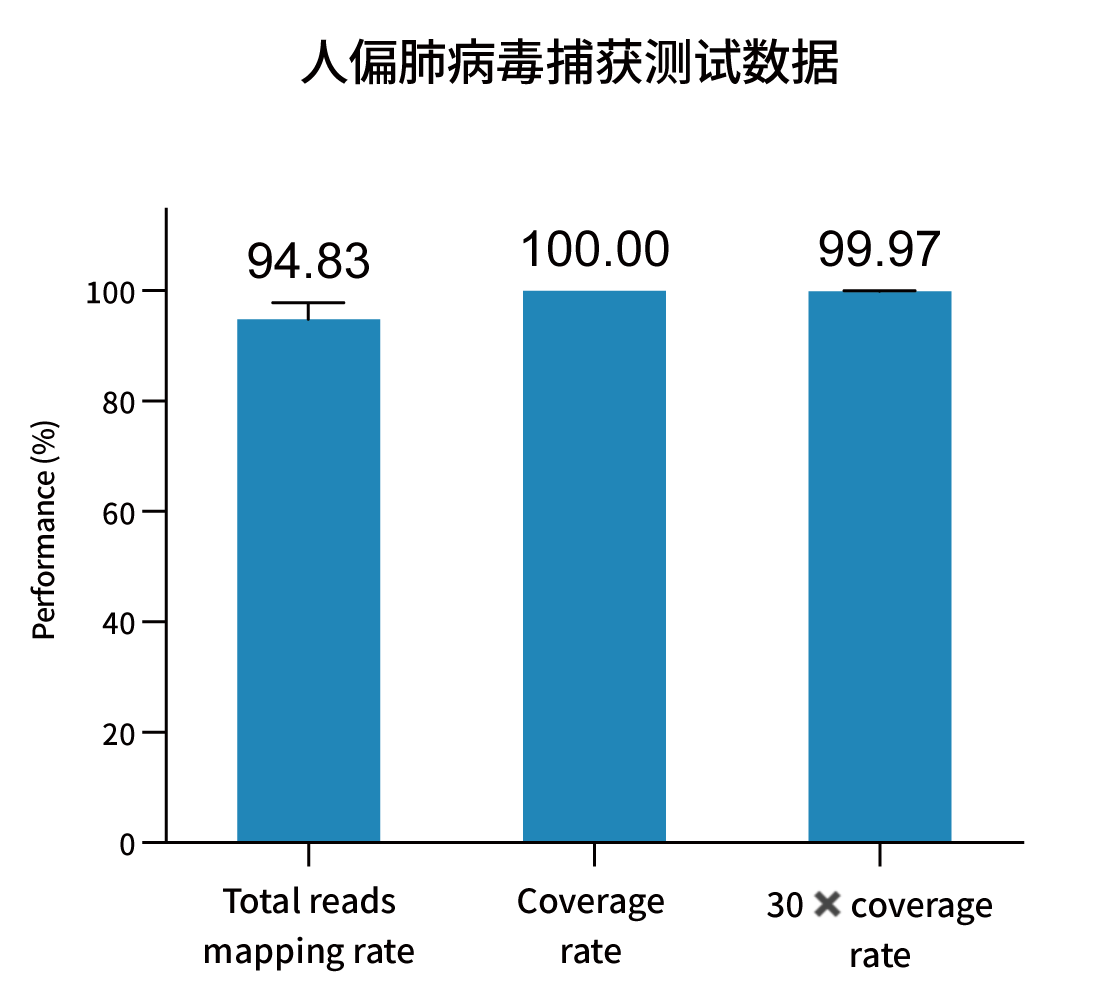
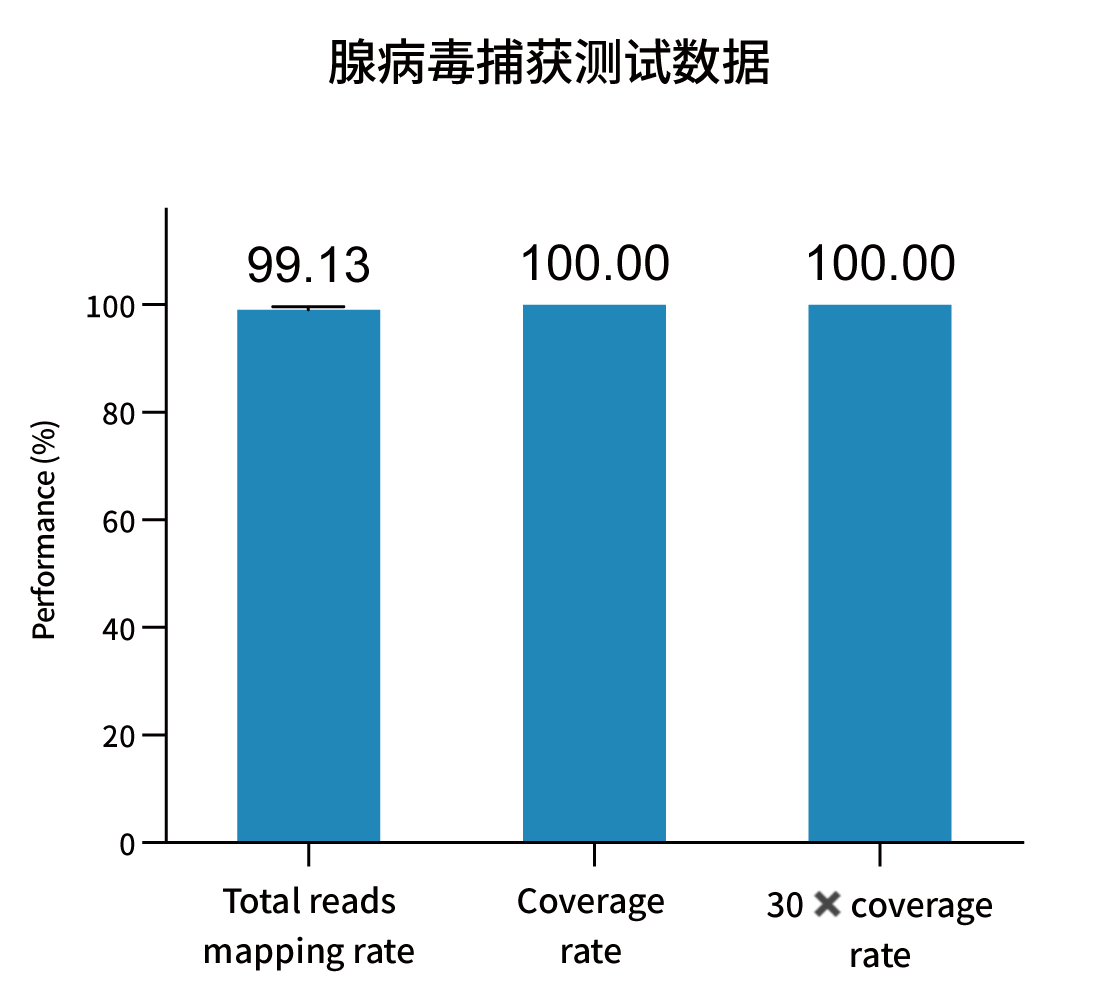
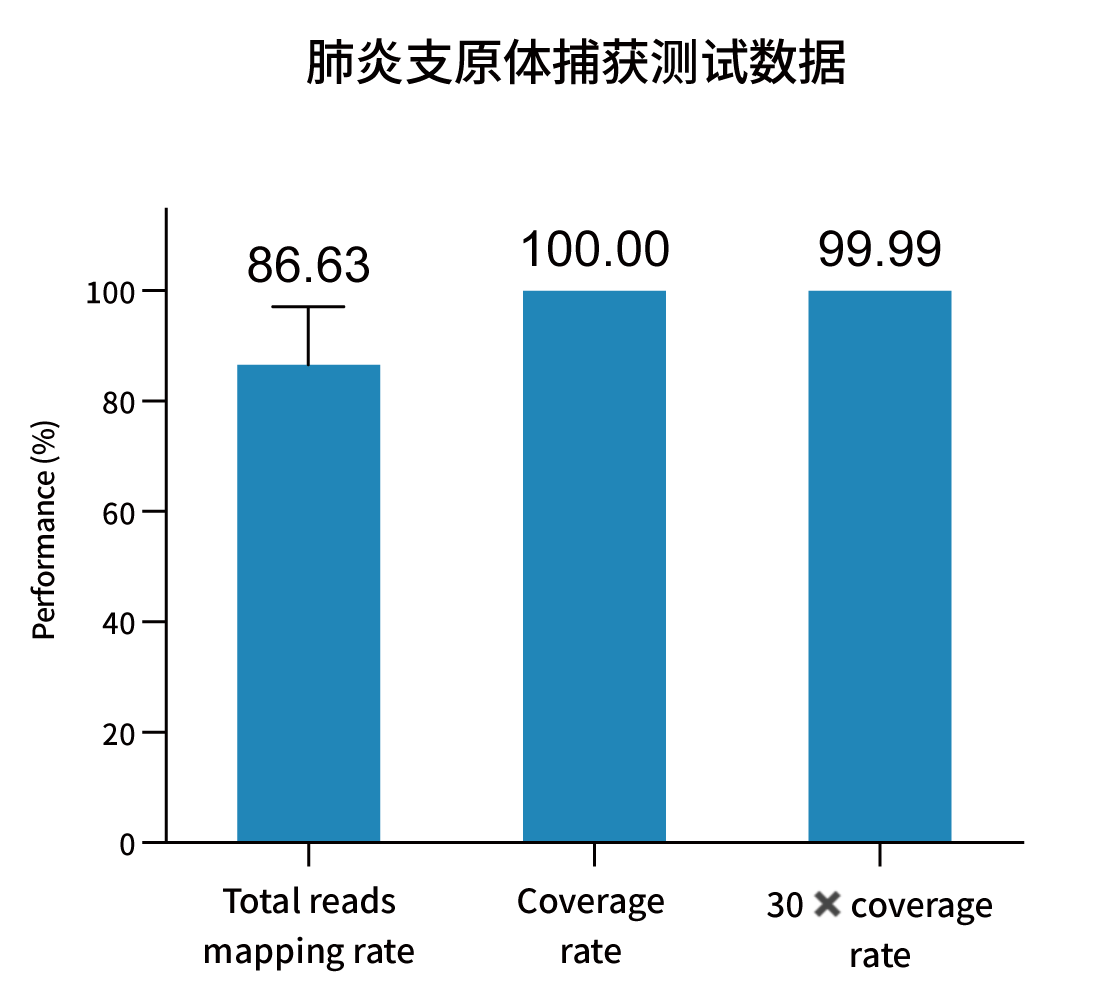
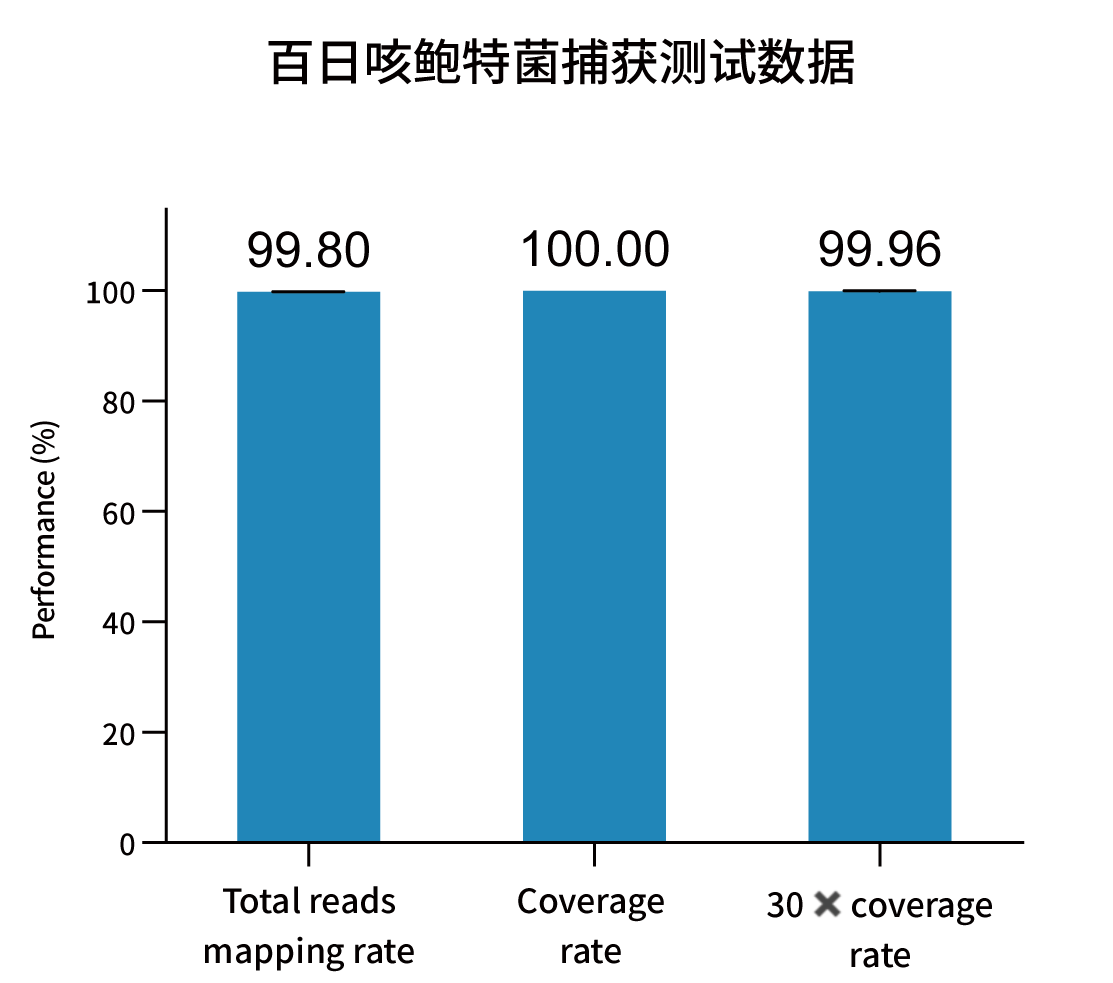
Ordering Information — Catalog & Pack Sizes
Available pack sizes and catalog numbers for each panel.
| Panel | 16 rxn | 96 rxn | Order |
|---|---|---|---|
| Influenza A/B/C Virus Panel | PH2000051 | PH2000052 | |
| SARS-CoV-2 Panel | PH2001721 | PH2001722 | |
| Pathogenic Coronavirus Panel | PH2001541 | PH2001542 | |
| Human Respiratory Syncytial Virus Panel | PH2000701 | PH2000702 | |
| Human Metapneumovirus Panel | PH2001131 | PH2001132 | |
| Human Parainfluenza Virus Panel | PH2001381 | PH2001382 | |
| Human Mastadenovirus Panel | PH2001061 | PH2001062 | |
| Rhinovirus Panel | PH2001881 | PH2001882 | |
| Varicella-Zoster Virus Panel | PH2004171 | PH2004172 | |
| Mycoplasma Pneumoniae Panel | PH2000031 | PH2000032 | |
| Chlamydia Psittaci Panel | PH2002161 | PH2002162 | |
| Chlamydia Pneumoniae Panel | PH2005771 | PH2005772 | |
| Chlamydia Trachomatis Panel | PH2006071 | PH2006072 | |
| Bordetella Pertussis Panel | PH2001121 | PH2001122 | |
| Mycobacterium Tuberculosis Panel | PH2001181 | PH2001182 | |
| Haemophilus Influenzae Panel | PH2003791 | PH2003792 | |
| Streptococcus Pyogenes Panel | PH2003471 | PH2003472 | |
| Klebsiella Pneumoniae Panel | PH2003601 | PH2003602 | |
| Acinetobacter Baumannii Panel | PH2003581 | PH2003582 | |
| Streptococcus Pneumoniae Panel | PH2003591 | PH2003592 |